|
Storage and computational demands strain the capabilities of even well-endowed research institutions’ high-performance compute (HPC) infrastructure - rendering the analysis of these datasets unaffordable using methods common in fields accustomed to smaller datasets (e.g. Though large-scale datasets present unique research opportunities, they also constitute immense challenges. This development is accompanied by a growing awareness of the importance to make the data more findable, accessible, interoperable, and reusable (FAIR) 2, and increasing availability of research standards and tools that facilitate data sharing and management 3. The Wind Integration National Dataset (WIND) Toolkit 1, CERN data (), or NASA Earth data () are only some of the prominent examples of large, openly shared datasets across scientific disciplines. The amount of data available to researchers has steadily grown, but over the past decade, a focus on diverse, representative samples has resulted in datasets of unprecedented size. 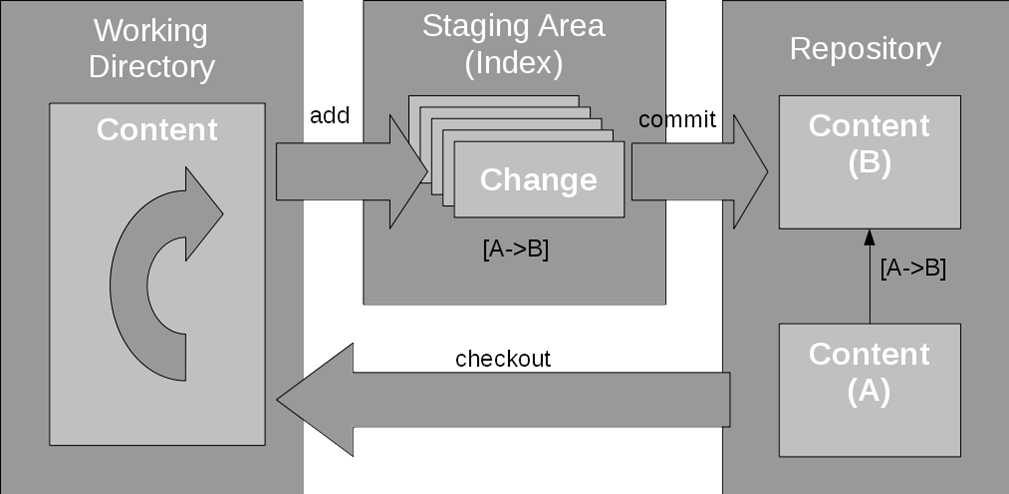
We demonstrate the framework’s performance using two showcases: one highlighting data sharing and transparency (using the dataset) and another highlighting scalability (using the largest public brain imaging dataset available: the UK Biobank dataset). It affords the capture of machine-actionable computational provenance records that can be used to retrace and verify the origins of research outcomes, as well as be re-executed independent of the original computing infrastructure. The framework attempts to minimize platform idiosyncrasies and performance-related complexities. Here we introduce a DataLad-based, domain-agnostic framework suitable for reproducible data processing in compliance with open science mandates. However, they also pose considerable challenges for the findability, accessibility, interoperability, and reusability (FAIR) of research outcomes due to infrastructure limitations, data usage constraints, or software license restrictions. Large-scale datasets present unique opportunities to perform scientific investigations with unprecedented breadth. The recipe used to build the CAT Singularity container 53 (10.5281/zenodo.6021002) is publicly available at. Īll scripts used to process the data 52 are publicly available at (10.5281/zenodo.6019782). The studyforrest derivatives computed by the tutorial workflow 51 (10.5281/zenodo.6019794) are publicly available from. Structural data from the Studyforrest project 46 (10.12751/g-node.zdwr8e) are available at. Interested parties can apply for data from UK Biobank directly, at. ĭata from the UK Biobank project were obtained from a third party, UK Biobank, upon application. 
Meyer K, Hanke M, Halchenko Y, Poldrack B, Wagner A.psychoinformatics-de/fairly-big-processing-workflow: Publication. psychoinformatics-de/fairly-big-processing-workflow-tutorial: Publication. Hanke M, Wagner AS, Waite LK, Mönch C.Hanke M, Waite LK, Poline J-B, Hutton A.Dvc: Data version control - git for data amp models. fMRIPrep: a robust preprocessing pipeline for functional MRI. 
The Brain Imaging Data Structure (BIDS) Specification.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
